Cancer Registry Pathology Report
Introduction
This is a draft of the Cancer Registry Content Profile (CRC) supplement to the PCC Technical Framework. This draft is a work in progress, not the official supplement or profile.
Profile Abstract
The Cancer Registry Content Profile (CRC)
This profile defines the implementation of HL7 2.3.1 to create a message from the completed pathology report and transmit it to the Cancer Reistry.
Glossary
Refer to the Glossary posted on CDC's National Program of Cancer Registries Modeling Electronic Reporting Project website: www.cdc.gov/cancer/npcr/informatics/merp/TBD
Issue Log
Open Issues
- Issue: Glossary: Use MERP glossary document or consolidated glossary tool?
- Refer to the existing document
- Issue: Dependencies: need help understanding this component
- Depends on Care Management with the V2 option (and by reference ATNA and CT)
- Issue: Under Profile Name: Add more information about the adverse effect of not getting full reporting.
- Issue: Options: need help understanding this component
- Won't have any.
- Issue: Groupings: Does this need to be modified for the CRC profile?
- Yes, not sure how
- Issue: Transaction Definitions: need help understanding this component
- Not relevant to this profile
- Issue: What information do we need to include in 3.1.2, 3.1.3, 3.2, 3.3, 3.4? The current Content Profile refers the reader to the NAACCR Standard already published.
- Just reference the NACCR Documentation on this section.
Closed Issues
Volume I
Add the following bullet to the list of profiles
- Cancer Registry Content - The Cancer Registry Content profile identifies the pathology data to be sent to Cancer Registries
Dependencies
Add the following row(s) to the list of dependencies
| Integration Profile | Dependency | Dependency Type | Purpose |
|---|---|---|---|
| Cancer Registry Content |
Profile Name
The Cancer Registry Content Profile (CRC)
Monitoring the occurence of cancer is a cornerstone of cancer control decision making. this monitoring, referred to as cancer surveillance, can be used to trigger case investigations, follow trends, evaluate the effectiveness of prevention measures such as screening and early detection programs and suggest public health priorities. Because most cancers are definitively diagnosed by microscopic examination of tissue, cancer surveillance programs rely n pathology reports to identify new cases and collect further information on cases previously reported.
Two challenges relate to pathology laboratory reporting of cancer cases. Laboratories may photocopy selected pathology reports and mail them to the cancer registry. This method is labor intensive and prone to missing cases because the laboratory staff is not aware of the full case definition for identifying a cancer case. Additionally, the cancer registry staff must re-enter the information into the cancer registry with a risk of data entry errors.
Alternately, laboratories may choose not to actively report cases, but allow cancer registry personnel to identify and photocopy appropriate pathology reports on site. This is very costly for the registry and has the same risk of data entry errors when the reports are manually entered into the registry database.
Use Cases
- Pre-condition
A woman goes in for a routine mammogram and a suspicious area is noted. The woman is referred to her family physician who does a biopsy and sends the tissue sample to the pathology laboratory for review and diagnosis.
The pathology laboratory receives the specimen, logs it into the laboratory information system (LIS) and prepares the specimen for analysis. The pathologist analyzes the specimen and dictates his/her findings, which are then transcribed into the LIS. The pathologist verifies the accuracy of the report and signs the transcribed report. The LIS marks the pathology report as "Final", triggering the use case events.
- Events
The LIS gathers information from its database and other appropriate databases to create a messages that contains all of the required data elements. The LIS validates the data to ensure it conforms to the HL7 2.3.1 ORU-RO1 specification and transmits it to the Cancer Registry using a secure connection.
The Cancer Registry receives the HL7 messages, parses it into the processing database and determines that a cancer diagnosis has been documented. The registrar codes the medical information. The pathology data and coded information is exported to the Cancer Registry Database and the message is archived.
- Post-condition
Pathology information has been added to the Cancer Registry Database.
The message is available for future use as needed.
The following diagram constrains the scope of the Content Profile.
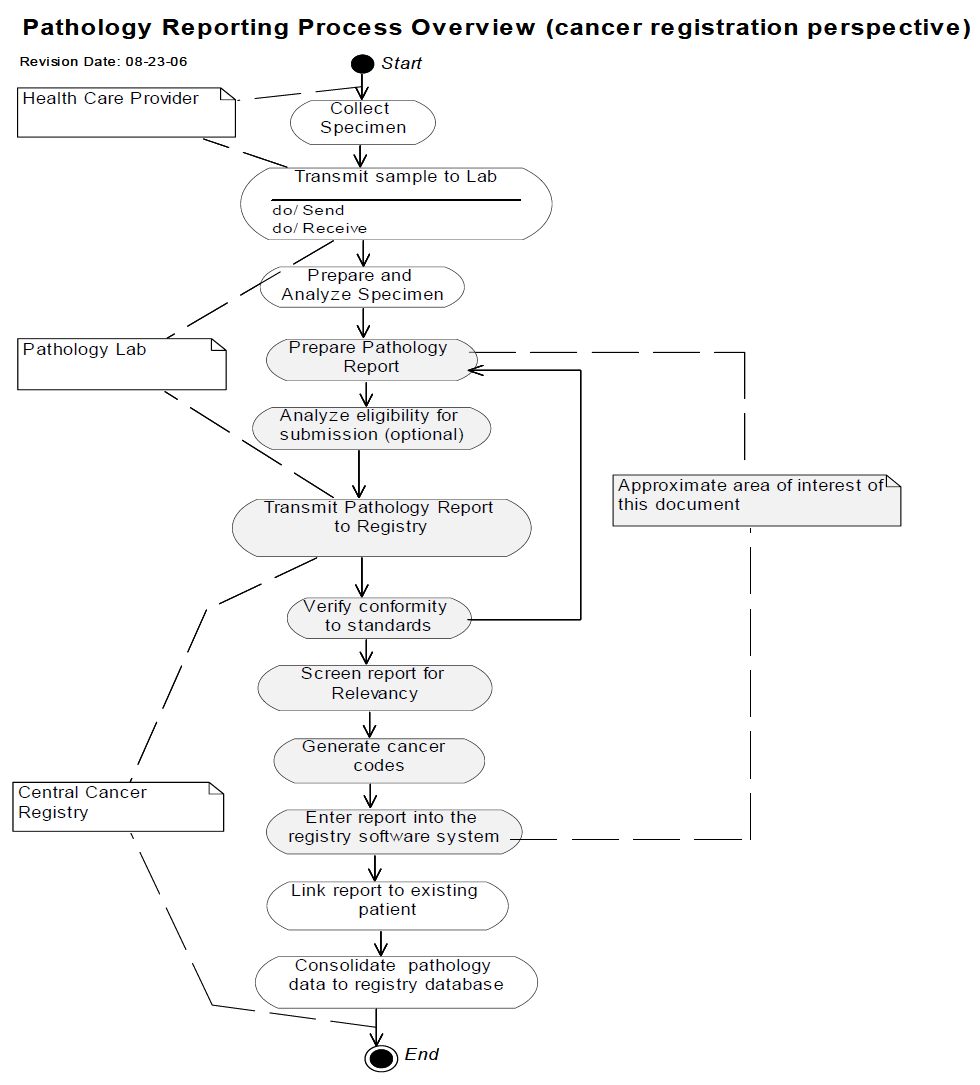
Actors/Transaction
There are two actors in this profile, the Content Creator and the Content Consumer. Content is created by a Content Creator and is to be consumed by a Content Consumer. The sharing or transmission of content from one actor to the other is addressed by the appropriate use of IHE profiles described below, and is out of scope of this profile. A Document Source or a Portable Media Creator may embody the Content Creator Actor. A Document Consumer, a Document Recipient or a Portable Media Importer may embody the Content Consumer Actor.
The sharing or transmission of content or updates from one actor to the other is addressed by the use of appropriate IHE profiles described by section 3.7 Content Bindings with XDS, XDM and XDR found in the Patient Care Coordination Technical Framework

Options
| Actor | Option | Section |
|---|---|---|
| Content Consumer | View Option (1) Document Import Option (1) |
PCC TF-1: 2.13.1 PCC TF-1: 2.13.2 |
| Content Creator | Referral Option (1) Discharge Summary Option (1) |
PCC TF-1: 2.13.5 PCC TF-1: 2.13.6 |
Note 1: The Actor shall support at least one of these options.
Grouping
Content Bindings with XDS, XDM and XDR
It is expected that the transfers of care will occur in an environment where the physician offices and hospitals have a coordinated infrastructure that serves the information sharing needs of this community of care. Several mechanisms are supported by IHE profiles:
- A registry/repository-based infrastructure is defined by the IHE Cross Enterprise Document Sharing (XDS) and other IHE Integration Profiles such as patient identification (PIX & PDQ), and notification of availability of documents (NAV).
- A media-based infrastructure is defined by the IHE Cross Enterprise Document Media Interchange (XDM) profile.
- A reliable messaging-based infrastructure is defined by the IHE Cross Enterprise Document Reliable Interchange (XDR) profile.
- All of these infrastructures support Security and privacy through the use of the Consistent Time (CT) and Audit Trail and Node Authentication (ATNA) profiles.
For more details on these profiles, see the IHE IT Infrastructure Technical Framework.
Content profiles may impose additional requirements on the transactions used when grouped with actors from other IHE Profiles.
Cross Enterprise Document Sharing, Media Interchange and Reliable Messages
Actors from the ITI XDS, XDM and XDR profiles embody the Content Creator and Content Consumer sharing function of this profile. A Content Creator or Content Consumer must be grouped with appropriate actors from the XDS, XDM or XDR profiles, and the metadata sent in the document sharing or interchange messages has specific relationships to the content of the clinical document described in the content profile.
Notification of Document Availability (NAV)
A Document Source should provide the capability to issue a Send Notification Transaction per the ITI Notification of Document Availability (NAV) Integration Profile in order to notify one or more Document Consumer(s) of the availability of one or more documents for retrieval. One of the Acknowledgement Request options may be used to request from a Document Consumer that an acknowledgement should be returned when it has received and processed the notification. A Document Consumer should provide the capability to receive a Receive Notification Transaction per the NAV Integration Profile in order to be notified by Document Sources of the availability of one or more documents for retrieval. The Send Acknowledgement option may be used to issue a Send Acknowledgement to a Document Source that the notification was received and processed.
Document Digital Signature (DSG)
When a Content Creator Actor needs to digitally sign a document in a submission set, it may support the Digital Signature (DSG) Content Profile as a Document Source. When a Content Consumer Actor needs to verify a Digital Signature, it may retrieve the digital signature document and may perform the verification against the signed document content.
Content Modules
Content modules describe the content of a payload found in an IHE transaction. Content profiles are transaction neutral. They do not have dependencies upon the transaction that they appear in.
Content Module 1
Process Flow
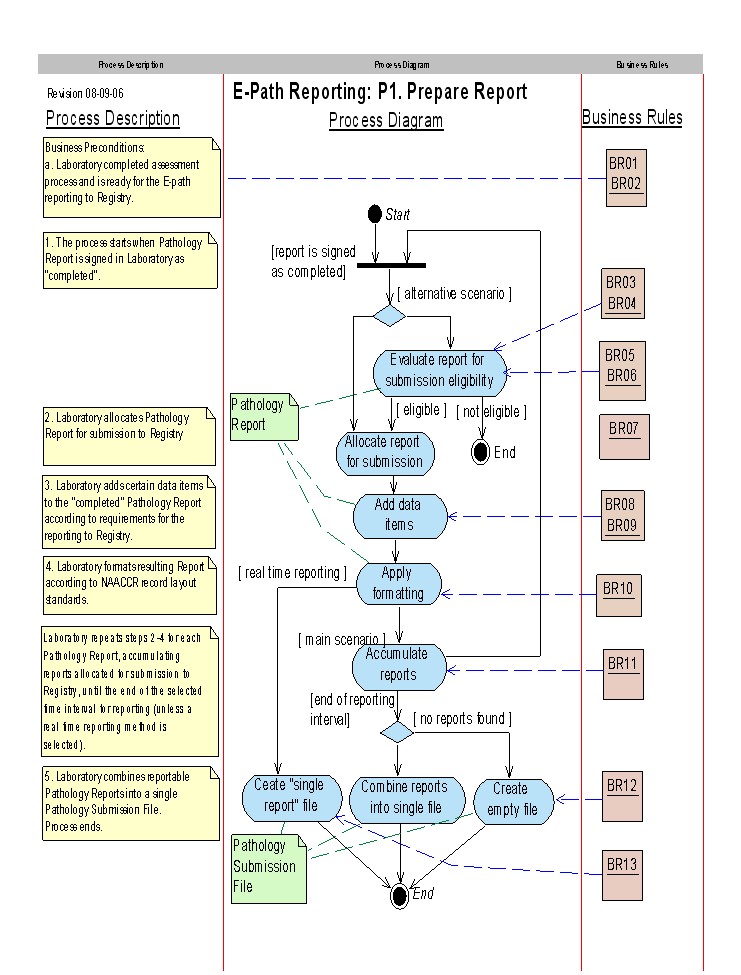
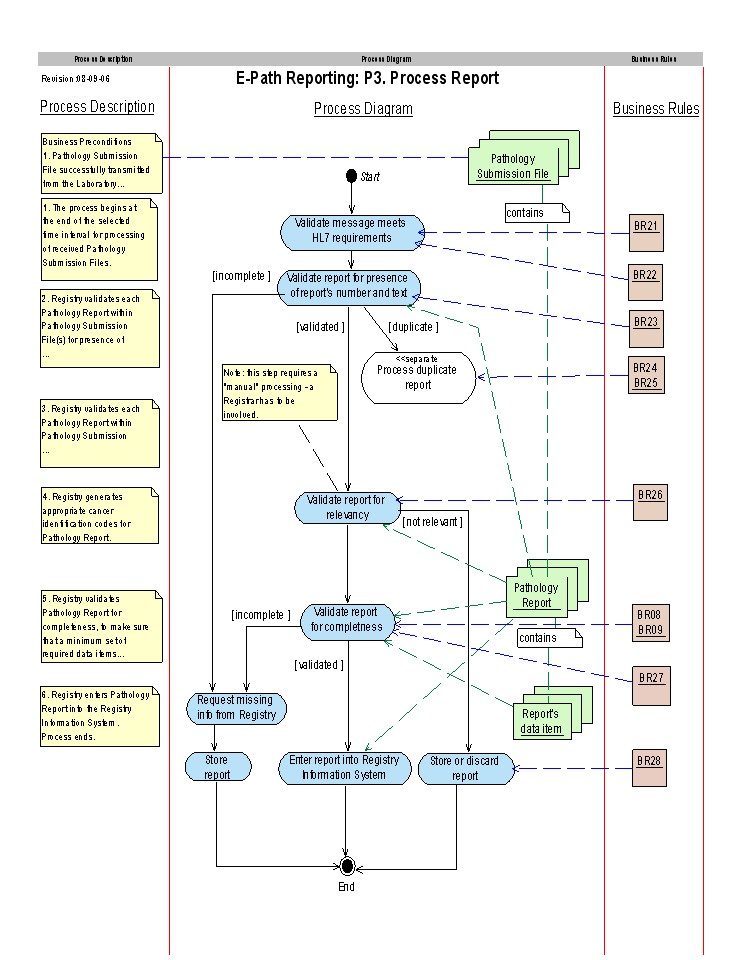
Refer to the North American Association of Central Cancer Registries Electronic Pathology (E-Path) Reporting Guidelines for a description of the business rules.
Actor Definitions
- Actor
The Content Creator is played by the Laboratory Information System.
The Content Consumer is played by the Cancer Registry Software System.
Transaction Definitions
- Transaction
- Definition
Volume II
Cancer Registry Content
Consists of the following documents:
Anatomic Pathology report
- histopathology
- flow cytometry
- cytology
- autopsy
- bone marrow
- peripheral smear
Standards
- HL7 2.3.1 ORU - Unsolicited Observation Message (event RO1)
- [1] : North American Association of Central Cancer Registries Standards for Cancer Registries Volume V: Pathology Laboratory Electronic Reporting; Version 2.1
- Standards for Identifying Pathology Reports
- NAACCR Search Term List
- [2]
- Case Definition
- NAACCR Standards for Cancer Registries Volume III: Standards for Completeness, Quality, Analysis, and Management of Data
- [3]
- Coding of Medical Information;
- International Classification of Diseases for Oncology (ICD-0) Third Edition
- World Health Organization
Data Element Index
Refer to NAACCR Standards for Cancer Registries Volume V: Pathology Laboratory Electronic Reporting Verison 2.1, Chapter 2.6
| Data Elements | Other Reference | LOINC Section or CDA Element |
|---|---|---|
| Data Element 1 | ||
| Data Element 2 | ||
| Data Element 3 |
Document Specification
| Data Element | Opt | Section | Template ID |
|---|---|---|---|
| Data Element 1 | R | 1.3.6.1.4.1.19376.1.5.3.1.3.X | |
| Data Element 2 | R2 | 1.3.6.1.4.1.19376.1.5.3.1.3.Y | |
| Data Element 3 | O | 1.3.6.1.4.1.19376.1.5.3.1.3.Z |
Section Template 1
| TemplateID | 1.3.6.1.4.1.19376.1.5.3.1.3.X | |
|---|---|---|
| Parent Template | 1.3.6.1.4.1.19376.1.5.3.1.3.Y | |
| General Description | This section shall ... | |
| LOINC Code | Opt | Description |
| #####-# | R | Description |
| Entries | Opt | Description |
| 1.3.6.1.4.1.19376.1.5.3.1.4.A | O | Description |
| Sub-sections | Opt | Description |
| 1.3.6.1.4.1.19376.1.5.3.1.3.D | R | Description |
Header Template 1
<entry>An XML Example</entry>
entry
Description of the entry element.
Entry Template 1
<entry>An XML Example</entry>
entry
Description of the entry element.