Enhanced Imaging Workflow IP: Difference between revisions
| Line 24: | Line 24: | ||
<li>Different scanning technologies can be used to digitize glass slides: bright field scanning, multispectral scanning, immunfluoreszence scanning, high resolution scanning (oil immersion). New scanning technologies will be developed. The application of these technologies depends on various factors and are influenced by specimen, and diagnostic question.</li> | <li>Different scanning technologies can be used to digitize glass slides: bright field scanning, multispectral scanning, immunfluoreszence scanning, high resolution scanning (oil immersion). New scanning technologies will be developed. The application of these technologies depends on various factors and are influenced by specimen, and diagnostic question.</li> | ||
</ol> | </ol> | ||
Related to image processing the following actors and transactions are described in the Technical Framework[http://www.ihe.net/Technical_Framework/index.cfm#pathology]: | |||
[[File:ActorsTransactions.png|800px|thumb|center|Transactions and Actors in the current TFAP]] | [[File:ActorsTransactions.png|800px|thumb|center|Transactions and Actors in the current TFAP]] | ||
<b>RAD-8 Modality Images Stored</b>: An Acquisition Modality sends acquired or generated images to the Image Archive. Defined in RAD TF-1:2.4, specified in RAD TF-2:4.8.<br> | |||
<b>RAD-10 Storage Commitment</b>: A requestor (Acquisition Modality or Evidence Creator) requests that the Image Manager confirm ownership for the specified DICOM objects (images, GSPS objects, Key Image Notes, Evidence Documents or any combination thereof) that the requestor stored in the Image Archive, thus allowing the sender to delete those objects now owned by the Image Manager. Defined in RAD TF-1:2.4, specified in RAD-TF-2:4.10.<br> | |||
<b>RAD-14 Query Images</b>: An Image Display queries the Image Archive for a list of entries representing images by patient, study, series, or instance. Defined in RAD TF-1:2.4, specified in RAD-TF-2:4.14.<br> | |||
<b>RAD-16 Retrieve Images</b>: An Image Display or an Imaging Document Consumer requests and retrieves a particular image or set of images from the Image Archive or an Imaging Document Source, respectively. Defined in RAD TF-1:2.4, specified in RAD-TF-2:4.16.<br> | |||
==3. Key Use Case== | ==3. Key Use Case== | ||
Revision as of 16:35, 23 August 2011
1. Proposed Workitem: Enhanced Imaging Workflow for Anatomic Pathology
- Editor: Thomas Schrader
The development of slide scanners offers the possibility to digitize glass slides in Pathology and this is the beginning of a new paradigm in this medical domain. New terms were created: Digital Pathology or Collaborative Pathology expressing the new potentials in the diagnostic process.
Not only digital patient data but also Whole Slide Images (WSI) are available now and can be used for
- support of diagnostic process
- analysis of images
- further analysis of clinical data.
With digitalization of glass slides two different discussions begun:
- How to integrate the digitalization process into the general diagnostic process in a Pathology Department?
- How to integrate efficiently image analysis into the diagnostic process?
Meanwhile different scanning technologies are avialable and can be used to evaluate different diagnostic questions e.g. multispectral images allow an enhancement in image quality and analysis.
--Tschrade 03:26, 17 August 2011 (CDT)
2. The Problem
At least two problems exist in integrating digialization and image analysis into the pathology processes:
- Image analysis can be done at various process steps during the diagnostic process and have different purposes: An explorative image analysis before the microscopic evaluation by the pathologist can extract regio of interests, enhance image quality or count some basic data such as lymph nodes. An analysis step after or during the microscopic evaluation is more specific: regio of interests can be marked very specifically. The results of such analysis can answer questions e.g. about mitosis rate, form factors of tumor cells, expression of tumor markers and much more.
- Different scanning technologies can be used to digitize glass slides: bright field scanning, multispectral scanning, immunfluoreszence scanning, high resolution scanning (oil immersion). New scanning technologies will be developed. The application of these technologies depends on various factors and are influenced by specimen, and diagnostic question.
Related to image processing the following actors and transactions are described in the Technical Framework[1]:
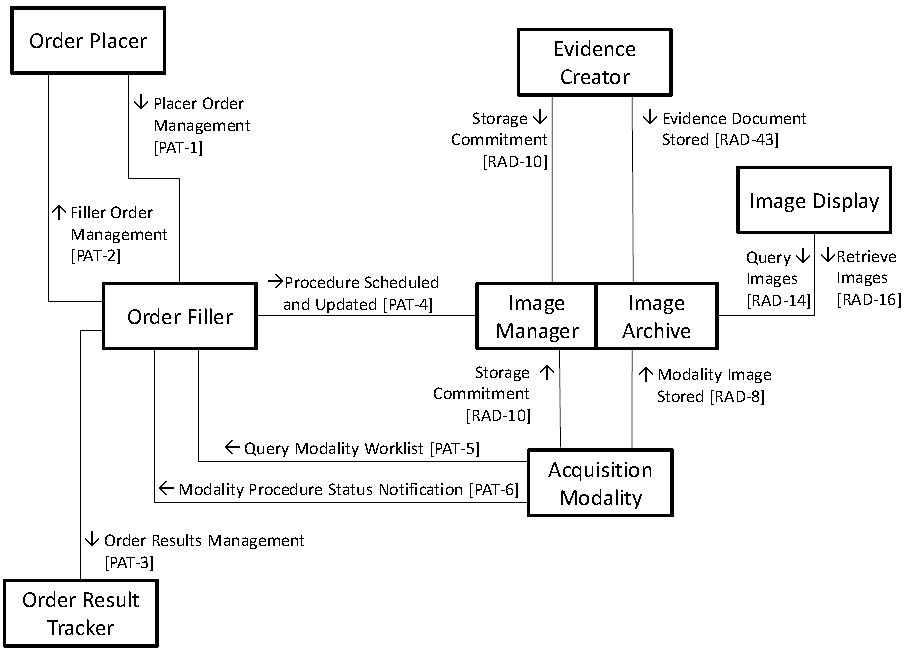
RAD-8 Modality Images Stored: An Acquisition Modality sends acquired or generated images to the Image Archive. Defined in RAD TF-1:2.4, specified in RAD TF-2:4.8.
RAD-10 Storage Commitment: A requestor (Acquisition Modality or Evidence Creator) requests that the Image Manager confirm ownership for the specified DICOM objects (images, GSPS objects, Key Image Notes, Evidence Documents or any combination thereof) that the requestor stored in the Image Archive, thus allowing the sender to delete those objects now owned by the Image Manager. Defined in RAD TF-1:2.4, specified in RAD-TF-2:4.10.
RAD-14 Query Images: An Image Display queries the Image Archive for a list of entries representing images by patient, study, series, or instance. Defined in RAD TF-1:2.4, specified in RAD-TF-2:4.14.
RAD-16 Retrieve Images: An Image Display or an Imaging Document Consumer requests and retrieves a particular image or set of images from the Image Archive or an Imaging Document Source, respectively. Defined in RAD TF-1:2.4, specified in RAD-TF-2:4.16.
3. Key Use Case
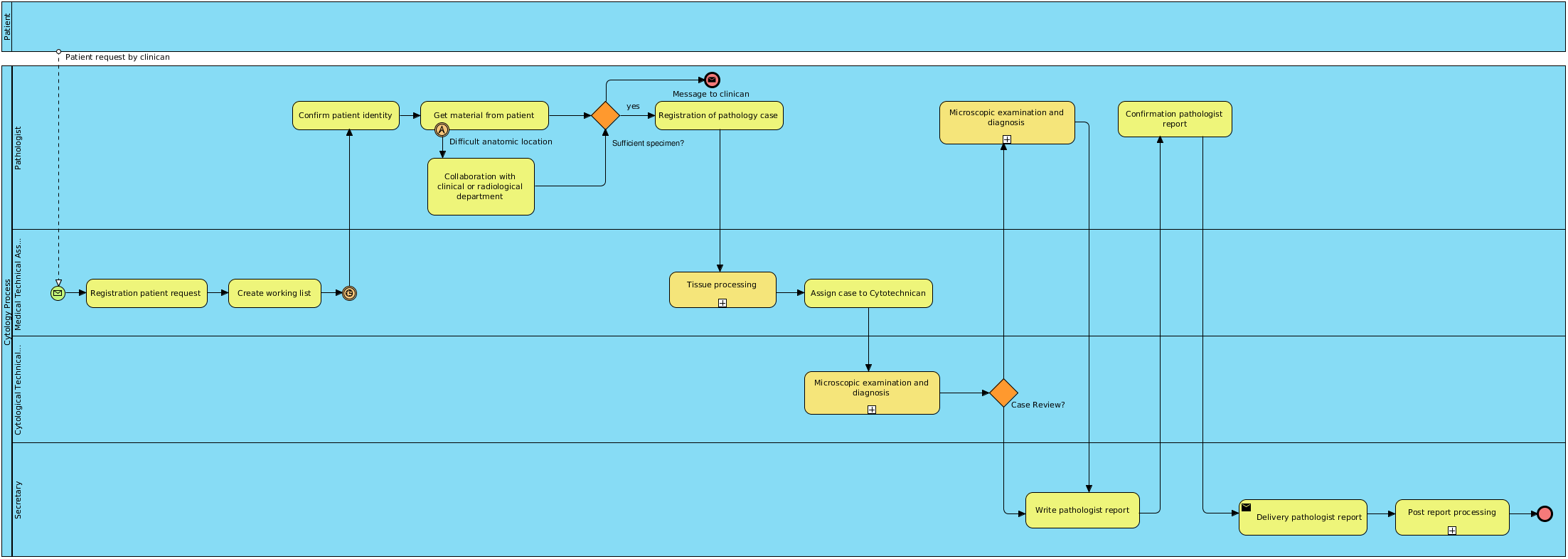
In the following figure the complete process of general pathology is presented. At the moment the digitalization of glass slide is not integrated in the routine processes.
The general diagnostic pathology process:

For cytology a slight different process was modeled:
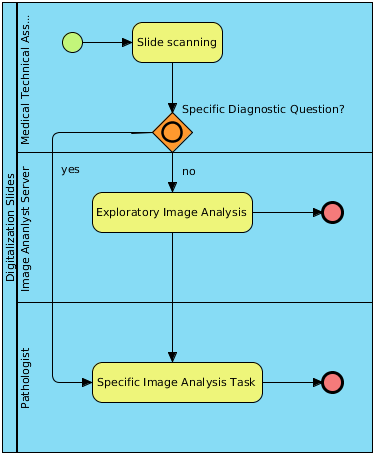
The used technology of scanning depends on various factors:
- Staining method: e.g. regulary staining, fluorescence staining
- Kind of specimen: e.g. high resolution (60x, 100x oil immersion) for cytology
- Analysis tasks
--TSchrader 15:17, 23 August 2011 (CDT)
4. Standards & Systems
5. Discussion
There is a clear need to integrate knowledge into the image processing in pathology.
New Actors in Pathology IHE Workflow
Two different variants are possible:
- Create a specialed (complete) actor for pathology workflow or
- Integrate existing actors from other TF's
Possible Actor from other TF
Treatment Management System (TMS) – An information system that manages oncology information and is responsible for the scheduling of radiotherapy activities (i.e. is a workflow manager). The TMS fulfils the role of a UPS-Pull ‘Worklist Manager’ SCP as described in DICOM Supplement 96 Part 17 Table Z.1-1. Note that in this profile the TMS fulfills the role of an Archive for the data objects, in which case the supplied AE Title in Input and Output Sequences is an AE Title managed by the TMS. Note that the TMS actors in other IHE-RO profiles have functionality that is different from that described in this profile. Technical Framework Supplement Treatment Delivery Workflow (TDW) [2]